Swati Bhattacharya
-
swaticb@che.iitb.ac.in

Courses
Research
Investigations of anti-HIV1 protein SAMHD1
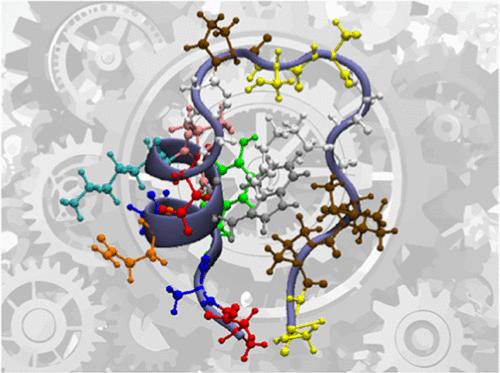
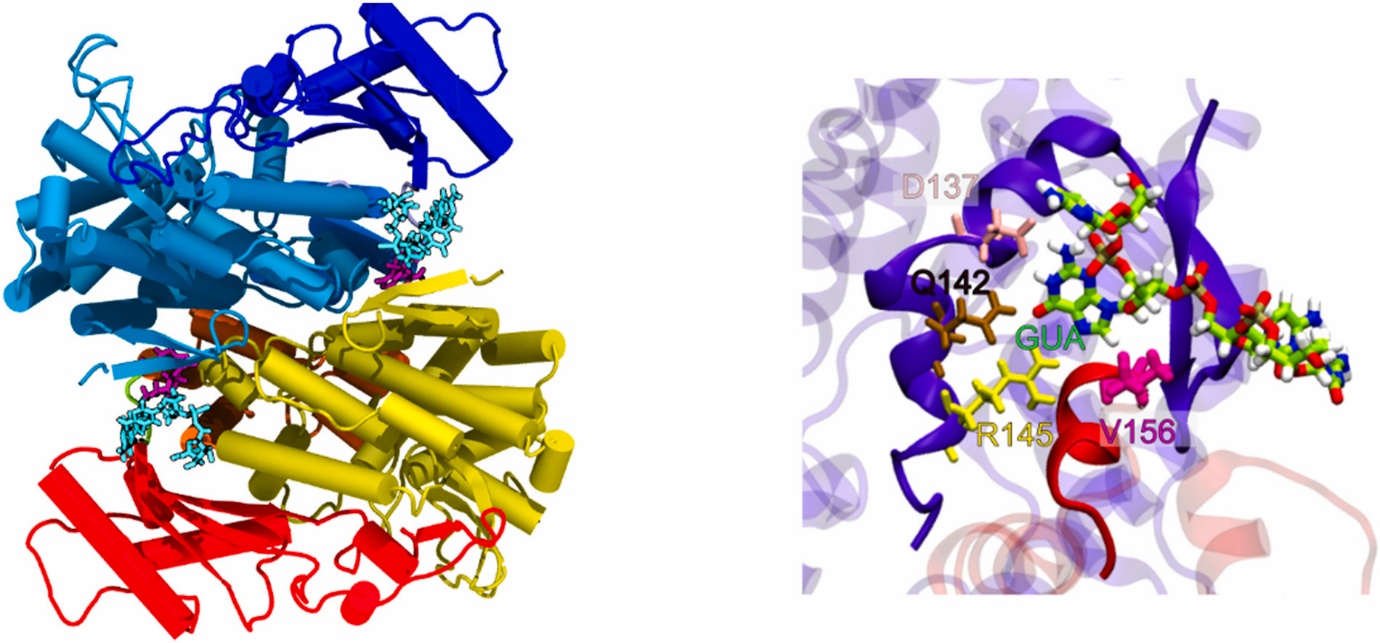
SAMHD1 (sterile alpha-motif and histidine aspartate domain containing protein-1) is a human protein that is responsible for the ineffective infection of myeloid cells and non-cycling CD4+ T cells by HIV-1, a feature attributed to its tri-phosphohydrolase activity; i.e, it is believed that by depleting the cellular pools of dNTPs it inhibits reverse transcription by retro-viruses. F There is increasing evidence of SAMHD1 playing a major role in oncogenesis. which may be exploited for cancer therapy. We have carried out a number of studies to uncover the working of SAMHD1 which can help to pave the way towards the development of a new class of anti-HIV drugs. We are currently engaged in using molecular dynamics simulations to get a mechanistic understanding of the allosteric regulation, phosphohydrolase activity and HIV-1 restriction activity. Our simulations have captured evidence of an exquisite reciprocal allosteric “handshake” occurring across monomeric units that plays a role not just in communicating signals between the allosite and catalytic sites of neighbouring monomers but also in transmitting signals from a regulatory site ( T592 cdk-1 phosphorylation site ) to the catalytic core.
G. Thapa, A. Bhattacharya and S. Bhattacharya, "Molecular dynamics investigation of DNA fragments bound to the anti-HIV protein SAMHD1 reveals alterations in allosteric communications ". J.Mol. Graphics and Model (2024). https://doi.org/10.1016/j.jmgm.2024.108748
G. Thapa, A. Bhattacharya and S. Bhattacharya, “Dimeric hold-states of anti-HIV-1 protein SAMHD1 are redox tunable.”, J Chem Inf Model. 60(12), 6377-6391,(2020). doi: 10.1021/acs.jcim.0c00629, (2020).
K.K. Patra, A. Bhattacharya and S. Bhattacharya. “Allosteric Signal transduction in HIV-1 Restriction Factor SAMHD1 proceeds via Reciprocal Handshake Across Monomers”, J. Chem. Inf. Modeling, 57(10), 2523-2538, (2017).
K. K. Patra, A. Bhattacharya and S. Bhattacharya. “Allosteric dynamics of SAMHD1 studied by molecular dynamics simulations” J. Phys.: Conf. Ser. 759 012026 (2016).
K.K. Patra, A. Bhattacharya and S. Bhattacharya. “Uncovering allostery and regulation in SAMHD1 through molecular dynamics simulations”. Proteins: Structure, Function and Bioinformatics, 85(7), 1266-1275 (2017). This was the cover article for the journal. https://doi.org/10.1002/prot.25287
Research Interests
Research Interests
The Bhattacharya research group is primarily engaged in the investigation of protein dynamics and mechanisms with applications to disease, intrinsic immunity, HIV therapeutics and enzymatic catalysis. The second thrust area of our research is the bottom-up modelling to unravel the long-timescale dynamics of proteins using theoretical and computational methods, primarily relying on molecular dynamics simulations.
Some of the milestones in our journey
- Uncertainty quantification of Markov state models and quantification of Validity time of MSMs
- Adaptive state-constrained MD
- Time-dependent Markov state models
- Local Structure and Dynamics Approach to quantify disorder in proteins
Methods Developed | Challenge Addressed | References |
Temperature Accelerated Markov State Modeling | Timescale gap due to high energy barriers |
|
Uncertainty in MSMs and quantification of Validity time | Missing States and Pathways | J. Chem. Phys. 143, 114109 (2015). |
Adaptive State-Constrained MD | Accelerate Extension of Validity time | J. Chem. Phys, 147, 152702 (2017) |
Time-dependent Markov State Model | Accelerate Extension of Validity time
| J. Chem. Phys, 147, 152702 (2017) |
Time-dependent Markov State Model | Non-equilibrium and driven systems | J. Chem Theory Comput., 13, 957-962 (2017) |
Accelerated Construction of kinetic networks using Steered MD | Use force-induced rapid transitions to build zero-force kinetic network models | J. Chem Theory Comput., 14, 5393-5405 (2018) |
Local Structure and Dynamics Approach to quantify disorder in proteins | Identify local independent moving parts, their conformations and kinetics from MD trajectories to build local kinetic networks | J. Phys. Chem. B, 128 (5), 1179-1187, (2024) |

Molecular Dynamics and the timescale problem
A major thrust area in our group is rare event acceleration techniques applied to biomolecular systems. The gap between timescales accessible to biomolecular simulations and many relevant biological processes presents a major obstacle to scientific discoveries in health and medicine using biomolecular simulations. We are developing a suite of techniques to combine rare event acceleration strategies such as using high temperature simulations3 or steered molecular dynamics with kinetic network model (Markov State Model or MSM) building techniques to overcome the timescale challenge. Our recent achievements in this area include uncertainty quantification of MSMs, development of time-dependent MSMs for non-equilibrium systems and accelerated construction of MSMs using SMD. Our long term goal in this regard is to reach realistic timescales for large proteins. In the future, we also plan to develop new software based on the methodologies developed that can be made publicly available.
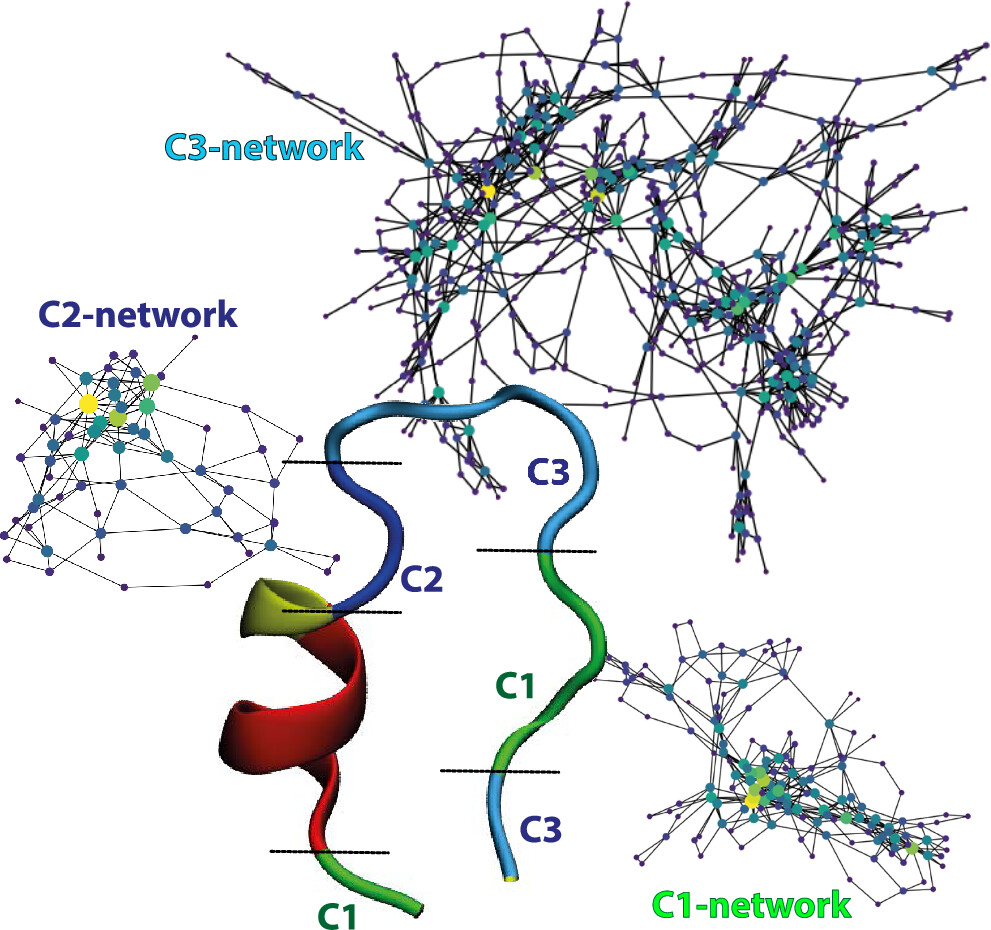
1.N. Fazeel, A. Chatterjee and S. Bhattacharya, "Quantifying Disorder in a Protein by Mapping its Locally Correlated Structure and Kinetics", J. Phys. Chem. B, 128 (5), 1179-1187, (2024). doi:https://doi.org/10.1021/acs.jpcb.3c06251
2. S. Ghosh, A. Chatterjee and S. Bhattacharya, “Time Dependent Markov State Models for Single Molecule Force Spectroscopy”, J. Chem Theory Comput., 13, 957-962 (2017).
3. A. Chatterjee and S. Bhattacharya. “Uncertainty in a Markov State model with missing states and rates: Application to a room temperature kinetic model obtained using high temperature molecular dynamics” J. Chem. Phys. 143, 114109 (2015).
4. K. K. Patra, A. Bhattacharya and S. Bhattacharya. “Allosteric dynamics of SAMHD1 studied by molecular dynamics simulations” J. Phys.: Conf. Ser. 759 012026 (2016).
5. A. Chatterjee and S. Bhattacharya. “Probing the energy landscape of alanine dipeptide and deca-alanine using temperature as a tunable parameter” J. Phys.: Conf. Ser. 759 012024 (2016).
Investigation of intrinsically disordered proteins
Although most proteins require a defined three-dimensional structure in order to carry out their function, there has been a growing recognition over the past decade of the importance of a class proteins that do not adopt a defined three-dimensional structure and are yet important for cellular function. Intrinsically disordered proteins, as these are termed, may contain collapsed-disorder, semi-collapsed disorder or extended disorder under physiological conditions or in vitro. Collapsed disorder consisting mainly of molten globules which are formed by hydrophobic collapse, have stable but dynamic secondary structure and flexible and dynamic side-chains. A part of our long term goals is to develop tools to unravel how the fine balance between order and disorder in such proteins can influence their activies. Our focus is on IDPs linked to diseases.
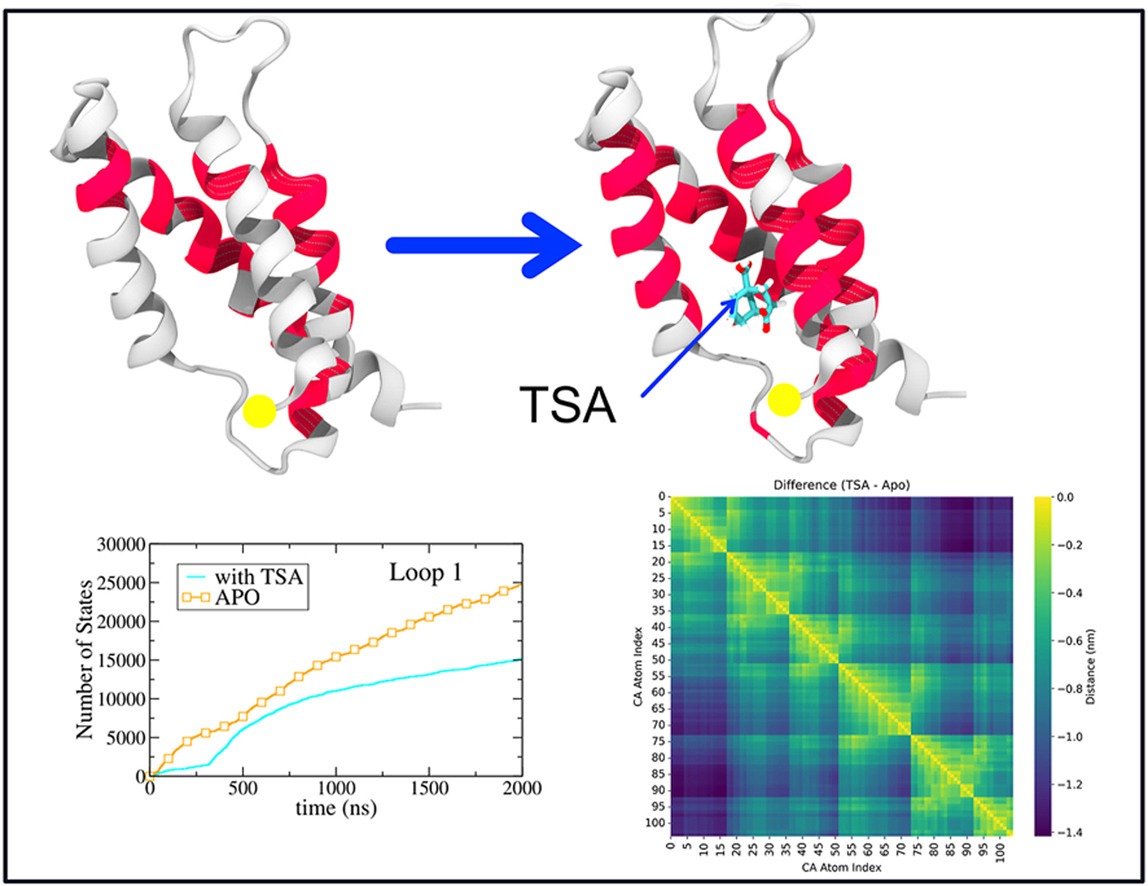
- F. Mahto, I. Hamid and S. Bhattacharya, “Molecular dynamics study of monomeric chorismate mutase shows large reduction in conformational diversity of loops upon binding of the transition state analog”, J.Mol. Graphics and Model 138, 109059 (2025) . https://doi.org/10.1016/j.jmgm.2025.109059
Research Areas
Sub Areas
- Molecular Simulations
